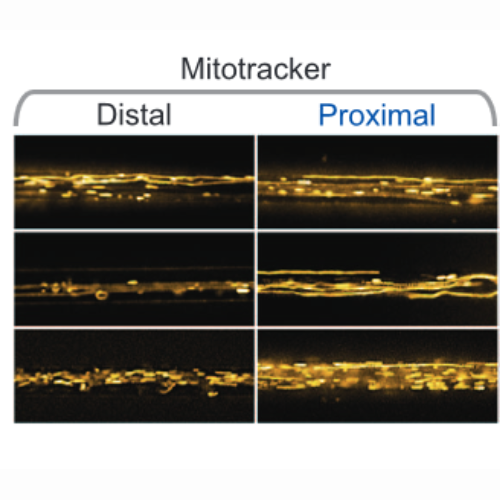
High content organelle trafficking enables disease state profiling as powerful tool for disease modelling.
Neurodegenerative diseases pose a complex field with various neuronal subtypes and distinct differentially affected intra-neuronal compartments. Modelling of neurodegeneration requires faithful in vitro separation of axons and dendrites, their distal and proximal compartments as well as organelle tracking with defined retrograde versus anterograde directionality. We use microfluidic chambers to achieve compartmentalization and established high throughput live organelle imaging at standardized distal and proximal axonal readout sites in iPSC-derived spinal motor neuron cultures from human amyotrophic lateral sclerosis patients to study trafficking phenotypes of potential disease relevance. Our semi-automated pipeline of organelle tracking with FIJI and KNIME yields quantitative, multiparametric high content phenotypic signatures of organelle morphology and their trafficking in axons. We provide here the resultant large datasets to enable systemic signature interrogations for comprehensive and predictive disease modelling, mechanistic dissection and secondary hit validation (e.g. drug screens, genetic screens). Due to the nearly complete coverage of analysed motility events, our quantitative method yields a bias-free statistical power superior over common analyses of a handful of manual kymographs.
Back to list